Team Simulates a Living Cell That Grows and Divides | Newswise
Newswise — By simulating the life cycle of a minimal bacterial cell — from DNA replication to protein translation to metabolism and cell division — scientists have opened a new frontier of computer vision into the essential processes of life.
The researchers, led by chemistry professor Zan Luthey-Schulten, present their findings in the journal Cell.
The team simulated a living cell at nanoscale resolution and recapitulated how every molecule within that cell behaved over the course of a full cell cycle. The work took many years, vast computer resources, large experimental datasets, a suite of experimental and computational techniques and an understanding of the roles, behaviors and physical interactions of thousands of molecular players. The researchers had to account for every gene, protein, RNA molecule and chemical reaction occurring within the cell to recreate the timing of cellular events. For example, their model had to accurately reflect the processes that allow the cell to double in size prior to cell division.
Watch a video of the full life cycle 4D simulation of a minimal bacterial cell.
To make the task more manageable, the team used a living “minimal cell” developed at the J. Craig Venter Institute in California. The version of the cell used in the new study, JCVI-syn3A — “Syn3A” for short — is a modified bacterium with a pared-down genome that carries only the genes needed to replicate its DNA, grow, divide and perform most of the other functions that make life possible.
“This is a three-dimensional, fully dynamic kinetic model of a living minimal cell that mimics what goes on in the actual cell,” Luthey-Schulten said. “Such a comprehensive undertaking was only possible through the combined efforts of a host of collaborators at the U of I as well as Harvard Medical School, where we systematically modeled the essential metabolism and other subcellular networks through a series of publications starting in 2018.”
The Syn3A cell has fewer than 500 genes, all of which reside on a single circular strand of DNA. The laboratories of study co-authors Angad Mehta, a professor of chemistry, and Taekjip Ha, of Boston Children’s Hospital and Harvard Medical School, generated additional experimental data that allowed the team to accurately simulate and validate numerous aspects of cell function.
Watch a video of the first-ever whole cell 4D simulation showing everything everywhere all at once.
“Most importantly, their work revealed the extent of DNA replication and that Syn3A’s cell division is symmetrical,” Luthey-Schulten said.
Both factors guided and validated the simulations performed by Zane Thornburg, a postdoctoral fellow at the Beckman Institute for Advanced Science and Technology and the Cancer Center at Illinois, and Andrew Maytin, a graduate student in Luthey-Schulten’s lab.
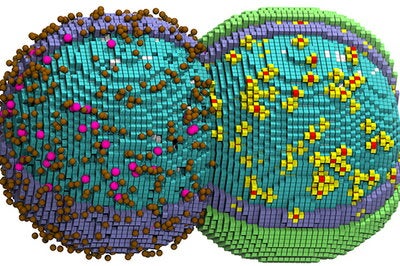
Like other bacterial cells, Syn3A has no nucleus. Every molecule that comprises and sustains it is either a component of its outer membrane, is transported into it from outside the cell or is assembled in the cytoplasm. The cell is so jam-packed with molecular players that, when creating high-resolution cartoons and animations of their computer simulations, the researchers had to render some of the components invisible. Making all the cellular proteins invisible, for example, allowed the scientists to see how Syn3A’s chromosome threads through the cell’s crowded interior.
Some processes were more computationally expensive than others, the team discovered. For example, Maytin realized that chromosome replication was slowing the whole simulation to a crawl, nearly doubling the time it took to capture the whole cell cycle. He determined that efficiently simulating the cell’s DNA replication process required its own dedicated graphics processing unit, while another GPU handled all other cellular dynamics. This allowed the team to simulate the full, 105-minute cell cycle in just six days of computer time.
Thornburg and Maytin struggled with the challenge of simulating cellular events occurring at the same time in various parts of the cell.
“I can’t overstate how hard it is to simulate things that are moving — and doing it in 3D for an entire cell was … triumphant,” Thornburg said. “One of the last big hurdles that Andrew and I had to solve was understanding how the membrane and the DNA talk to one another when both are moving.”
While the simulated cell cycle has its limitations — this was not an atom-by-atom simulation but instead averaged the dynamics of individual molecules — it yielded a surprisingly accurate accounting of the timing of cellular processes. In repeated simulations involving individual cells with slightly varying start conditions, the simulated cell cycle occurred, on average, within two minutes of the real-world cell cycle, Thornburg said. The work was repeatedly guided and tested against actual experimental outcomes, a process that allowed the scientists to refine their simulations.
The ability to accurately capture the ever-changing conditions within a living cell opens a new window on the foundations of living systems, Luthey-Schulten said.
“We have a whole-cell model that predicts many cellular properties simultaneously,” she said. “If you want to know what’s going on, say, in nucleotide metabolism, you can also look at what’s going on in DNA replication and the biogenesis of ribosomes. So the simulations can give you the results of hundreds of experiments simultaneously.”
Study co-authors also include Illinois chemistry alumnus Benjamin Gilbert and John Glass, who leads the J. Craig Venter Institute Synthetic Biology Group.
This work was conducted in the National Science Foundation’s Science and Technology Center for Quantitative Cell Biology at the U of I. Luthey-Schulten also is a professor of physics and a professor in the Beckman Institute at the U. of I. The research was conducted using the Delta advanced computing and data resource, which is supported by the NSF and the state of Illinois. Delta is a joint effort of the U of I and its National Center for Supercomputing Applications.